Istvan Botos, Ph.D.

Professional Experience
- Staff Scientist, NIDDK, NIH, 2005–Present
- Postdoctoral Fellow, Frederick Cancer Research and Development Center, NCI, NIH, 1999–2005
- Research Fellow, Hungarian Academy of Sciences, Institute of Biophysics, 1990–1992
- Ph.D., Texas A&M University, 1999
- M.Sc., University of Bucharest, Department of Biology, 1990
Research Goal
Understanding the molecular structure of integral membrane proteins from gram-negative bacteria is essential for designing alternative drugs against multi-drug resistant pathogenic bacteria.
Current Research
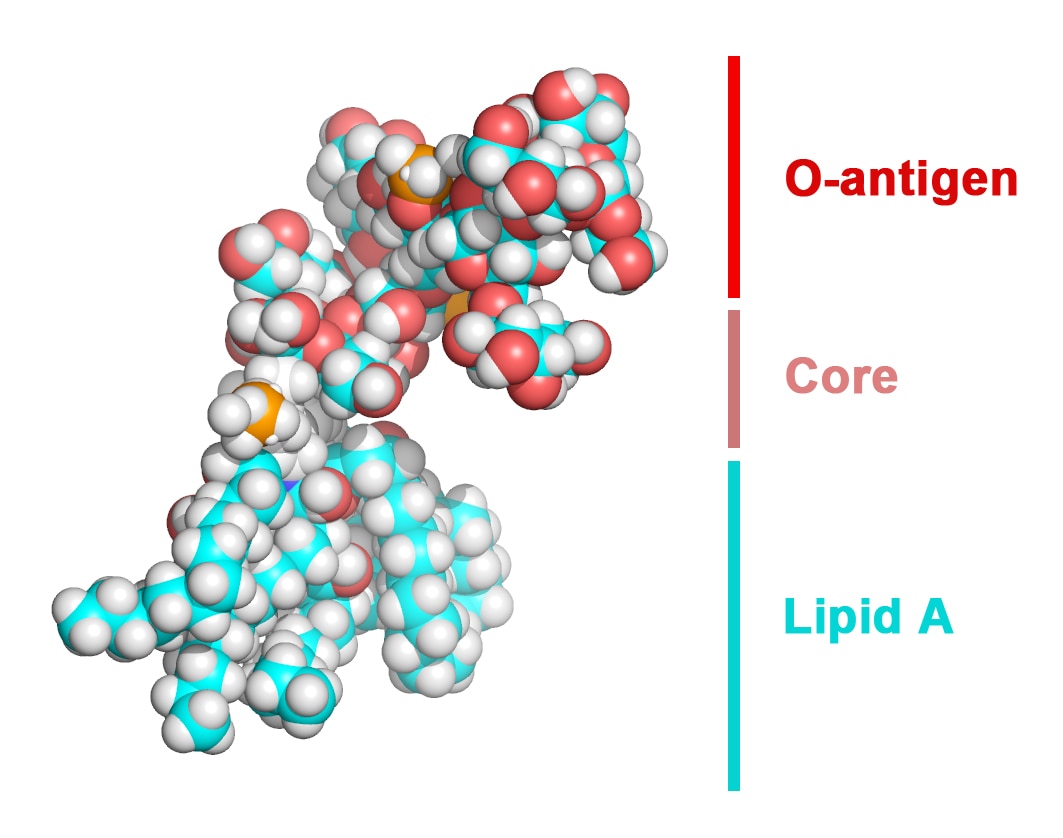
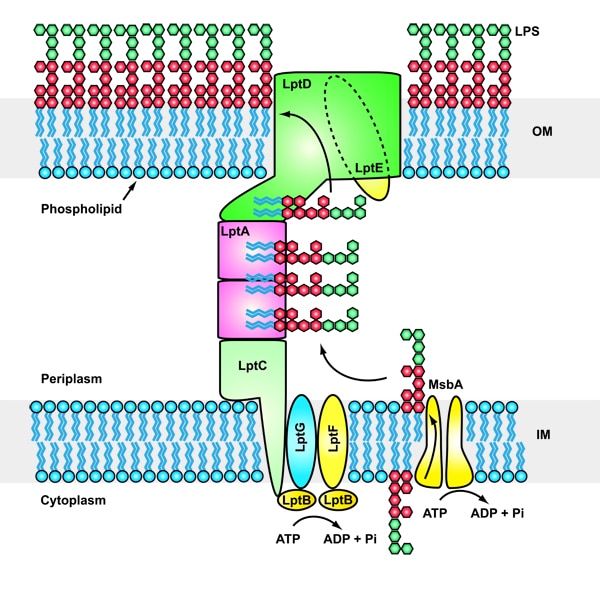
Gram-negative bacteria are protected from environmental stresses and toxins by a cell envelope consisting of an inner membrane, a periplasmic space, and an outer membrane. The outer membrane is asymmetric, with phospholipids in the inner leaflet and lipopolysaccharide (LPS) in the outer leaflet. LPS is essential for viability and is inserted into the membrane by a two-protein complex called LptDE. The complex is part of a larger protein machinery that transports LPS from the inner membrane to the outer membrane.
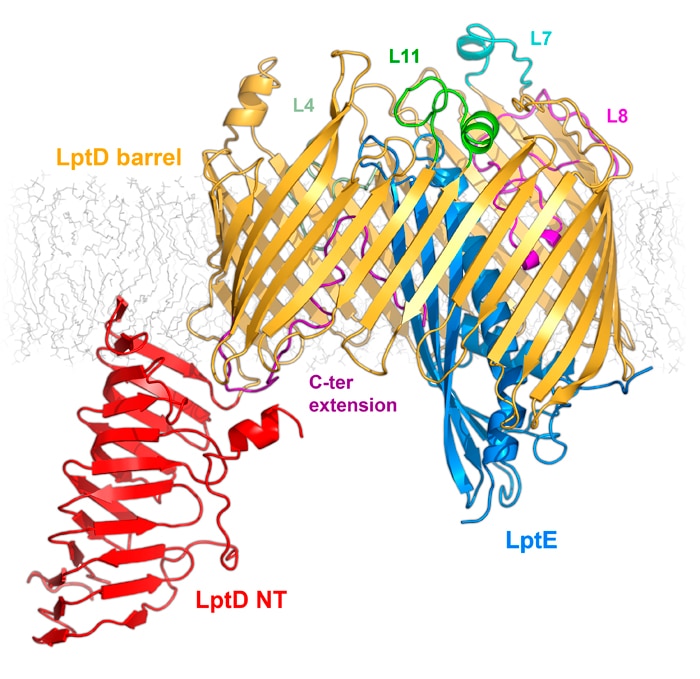
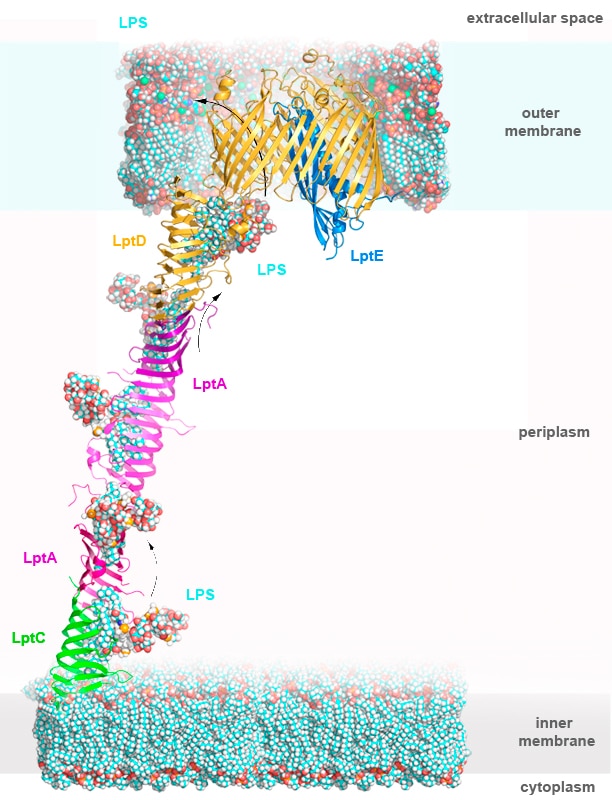
Our research group focuses on understanding integral membrane proteins. We primarily use X-ray crystallography and cryo-electron microscopy, which are methods for studying molecular structures at atomic resolution. We also conduct functional studies on these proteins. In particular, we determined crystal structures of the LptDE protein complexes from the pathogenic bacteria: Yersinia pestis, Klebsiella pneumoniae, and Pseudomonas aeruginosa. These membrane protein structures resemble a hollow barrel with a plug domain in the middle. From these structures we gained valuable insights into the transport mechanism of the LPS molecule. In the current model of the transport, a newly synthesized LPS molecule is transported from the inner membrane to the outer membrane by a slide-like bridge formed by several proteins of the pathway. This periplasmic bridge ends with the LptDE complex, which translocates the LPS molecule into the outer membrane.
Applying our Research
Because LPS export into the outer membrane is essential for bacterial viability, this process represents an excellent drug target in pathogenic bacteria.
Need for Further Study
The molecular structure and mechanism of action of some of the protein components in the LPS pathway are still unknown. Understanding the detailed mechanism of the components can be used to design specific molecules that block the pathway. Some of these molecules can be used to develop new therapies against multi-drug resistant pathogenic bacteria.
Select Publications
- The Name Is Barrel, β-Barrel.
- Hayashi S, Buchanan SK, Botos I.
- Methods Mol Biol (2024) 2778:1-30. Abstract/Full Text
- Variation of Tetrahydrofurans in Thyclotides Enhances Oligonucleotide Binding and Cellular Uptake of Peptide Nucleic Acids.
- Zheng H, Clausse V, Amarasekara H, Mazur SJ, Botos I, Appella DH.
- JACS Au (2023 Jul 24) 3:1952-1964. Abstract/Full Text
Research in Plain Language
Our research group focuses on understanding membrane proteins that are located on the surface of bacteria. We primarily use X-ray crystallography and cryo-electron microscopy, which are methods for studying molecular structures in detail. We also conduct functional studies. Understanding the structure of these proteins from pathogenic bacteria makes possible the design of new therapies against multi-drug resistant pathogenic bacteria.
Specifically, we determined the structures of protein complexes that transport LPS to the outer membrane of bacteria. LPS (or lipopolysaccharide) is a major component of the bacterial outer membrane and vital for the survival of pathogenic bacteria. Understanding the detailed mechanism of this transport process will lead to novel therapeutics against infectious disease.
Related Links
Research Images