Robert Craigie, Ph.D.

Professional Experience
- Ph.D., London, 1982
- B.S., London, 1978
Current Research
Our research focuses on mechanistic aspects of HIV DNA integration, which is an essential step in the replication of HIV and a frontline target of current drug regimens used to treat HIV/AIDS. We use biochemical approaches and cryo-electron microscopy to study the nucleoprotein complexes (intasomes) that mediate DNA integration, the molecular mechanism by which drugs block DNA integration, and how HIV can escape by acquiring mutations in the viral integrase enzyme.
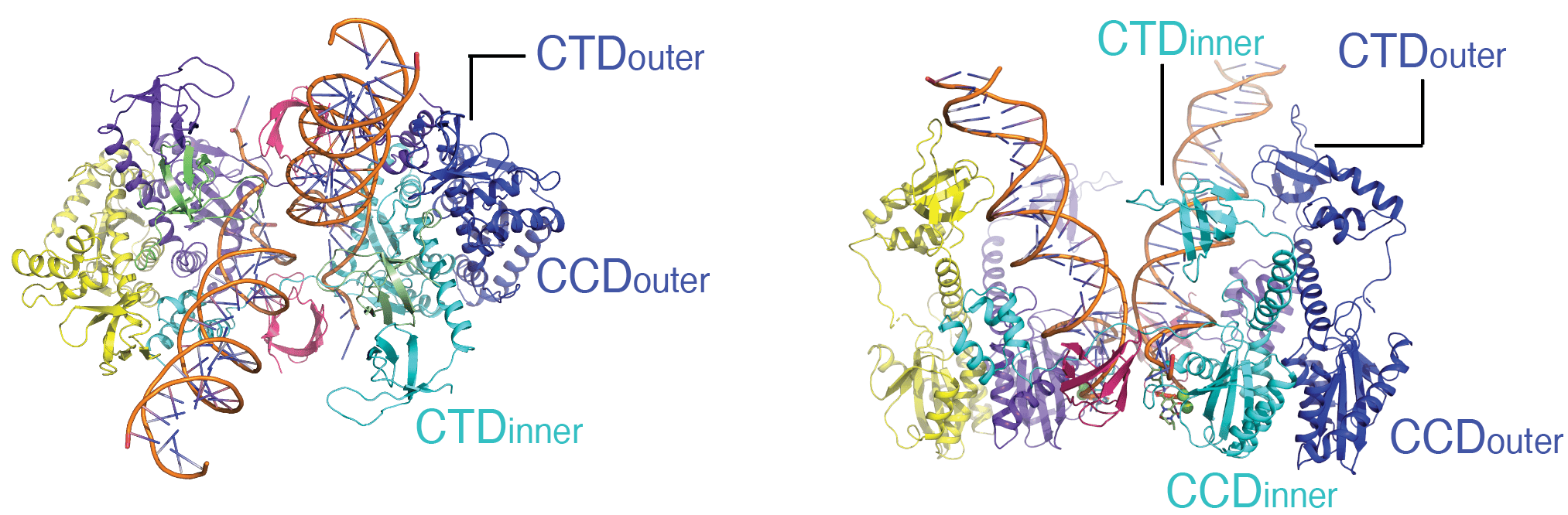
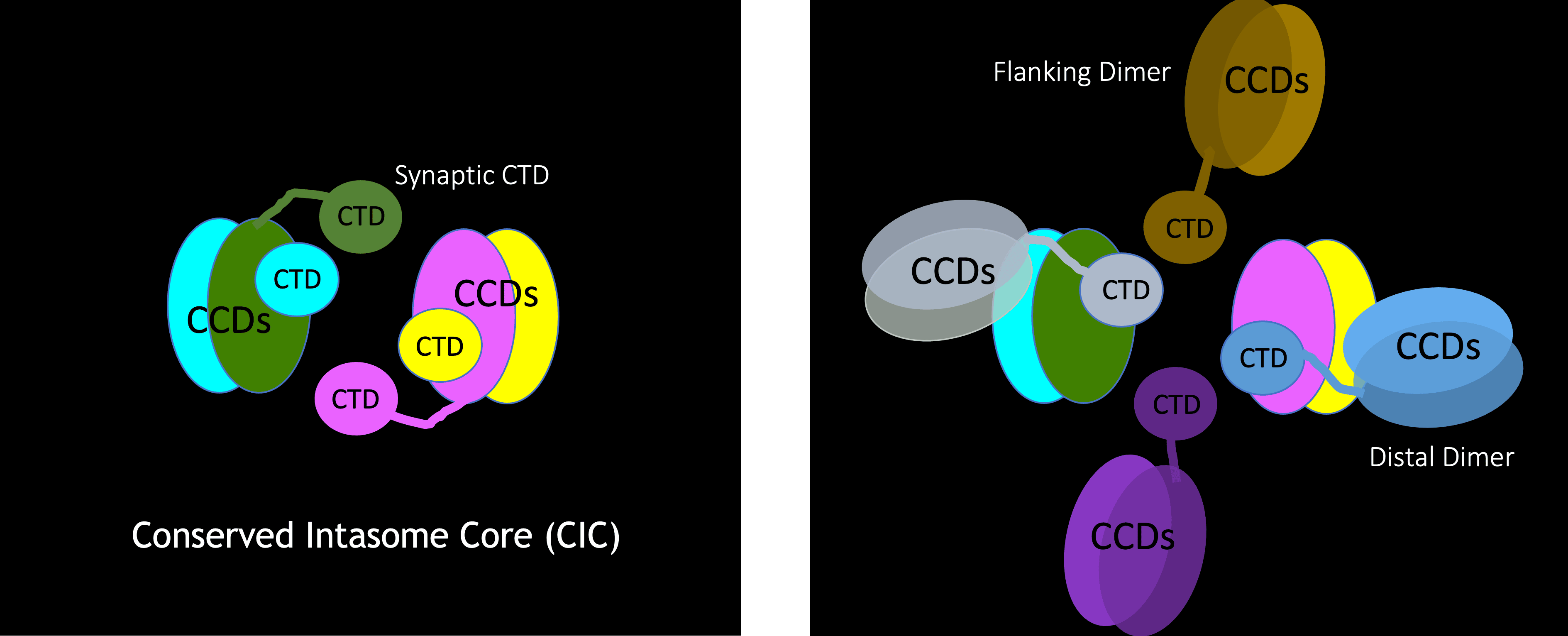
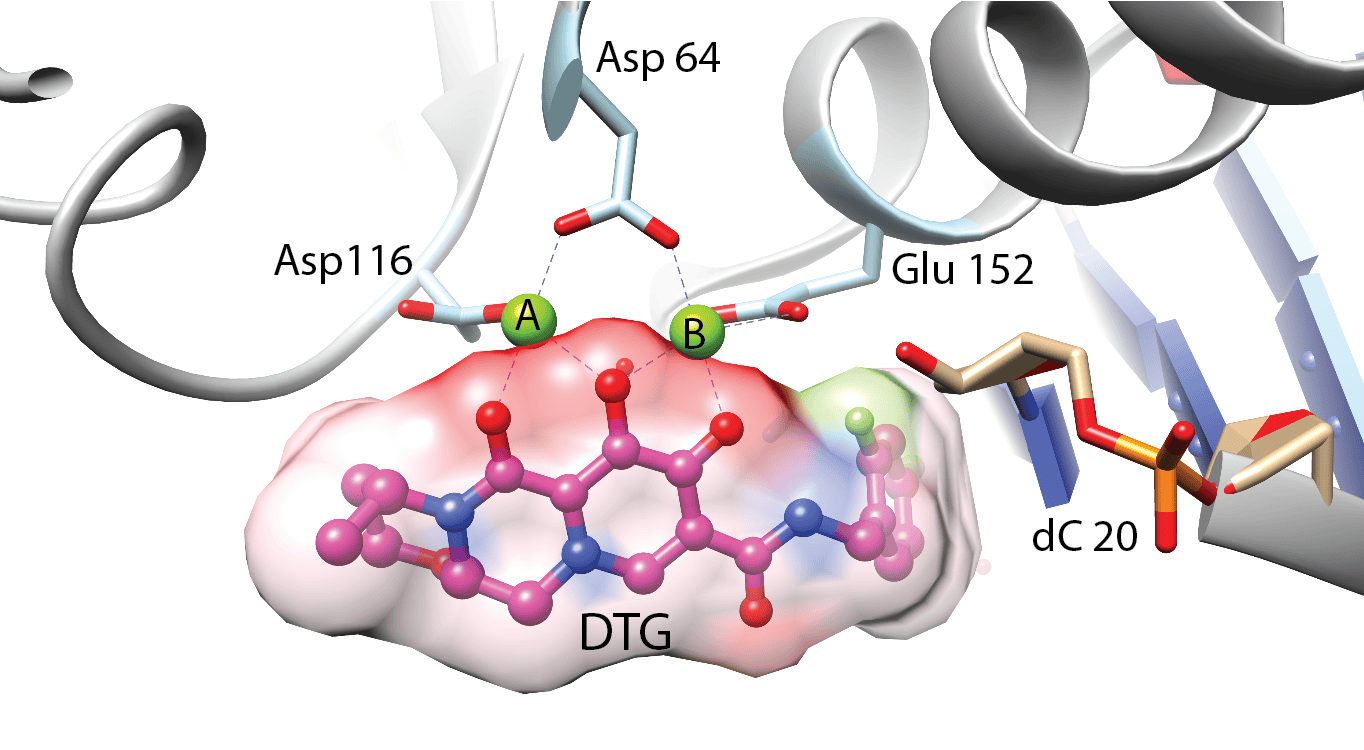
The key player in the retroviral DNA integration process is the virally encoded integrase protein. Integrase processes the ends of the viral DNA and covalently inserts these processed ends into host DNA. These reactions are catalyzed within a nucleoprotein complex between integrase and viral DNA ends called the intasome. We previously developed in vitro DNA integration reaction systems using purified HIV integrase, synthetic viral DNA ends and a divalent metal ion. These in vitro systems enabled the biochemical mechanism of integration to be elucidated. Drugs that inhibit HIV DNA integration target intasomes, rather than free integrase protein. High-resolution structures of HIV-1 intasomes are therefore needed to understand how these drugs act and how the virus can escape by evolving resistance mutations in integrase. Intasomes are present in only a few copies per infected cells, so the biochemical and biophysical studies that can be carried out with intasomes from infected cells are extremely limited. We have therefore devised methodologies to assemble intasomes from purified integrase and oligonucleotide DNA that mimics the viral DNA ends. Aggregation of HIV intasomes in vitro presented a major challenge to structural studies. Biochemical strategies and advances in cryo-electron microscopy have enabled these obstacles to be overcome. We are now able to routinely obtain structures of HIV intasomes in to 2-3 Å range of resolution. These structures have revealed a positionally conserved set of integrase domains around the viral and target DNA (the conserved intasome core), which is conserved in all intasome structures to date. These domains can be contributed by different protomers within the intasome. High resolution structures have been obtained for HIV intasomes in complex with the integrase inhibitors, such as dolutegravir, that are in clinical use. These structures define their mechanism of action and provide information that can leads to the design of improved derivatives; all the currently approved drugs were developed from lead compounds identified by high-throughput screening in the absence of structural information. Recent efforts have focused on structural studies of intasomes carrying resistance mutations in integrase in complex inhibitors. These structures reveal how mutations lead to drug resistance and suggest strategies for making escape by mutation more difficult.
Select Publications
- HIV-1 Integrase Assembles Multiple Species of Stable Synaptic Complex Intasomes That Are Active for Concerted DNA Integration In vitro.
- Li M, Yang R, Chen X, Wang H, Ghirlando R, Dimitriadis EK, Craigie R.
- J Mol Biol (2024 May 15) 436:168557. Abstract/Full Text
- Structural basis for strand-transfer inhibitor binding to HIV intasomes.
- Passos DO, Li M, Jóźwik IK, Zhao XZ, Santos-Martins D, Yang R, Smith SJ, Jeon Y, Forli S, Hughes SH, Burke TR Jr, Craigie R, Lyumkis D.
- Science (2020 Feb 14) 367:810-814. Abstract/Full Text
Research in Plain Language
We use biochemical, biophysical, and structural techniques such as cryo-electron microscopy to improve our understanding of how one type of virus, the retrovirus, proliferates and thrives. HIV is an example of a retrovirus. Unlike most living things, viruses do not encode everything they need to replicate. They must infect cells and depend, in part, on cellular machinery to make new copies of themselves.
DNA contains the instructions needed for an organism to develop, survive, and reproduce. To carry out these functions, DNA sequences must be converted into messages that can be used to produce proteins. In most viruses, DNA is transcribed to ribonucleic acid (RNA), and then RNA is translated into protein. Retroviruses function differently. The RNA of retroviruses is reverse transcribed into DNA, which is integrated into the host cell’s genome and then undergoes the usual transcription and translational processes to express the genes carried by the virus.
We focus especially on the role of one key protein: integrase. By what means does it process the ends of the viral DNA and insert them into host DNA? How can this process be interrupted? How do drugs that block HIV-1 replication by inhibiting integrase work? How can HIV-1 escape drugs by evolving resistance and how might improved drugs be developed?
Research Images