Irina V. Gopich, Ph.D.

Professional Experience
- Staff Scientist, NIDDK, NIH, 2004–Present
- Visiting Fellow, NIH, 1999–2004
- Visiting Fellow, Hebrew University of Jerusalem, 1997, 1999
- Ph.D., Institute of Chemical Kinetics and Combustion, 1996
Research Goal
Our research is focused on understanding the physical processes that play key roles in the activity and function of biomacromolecules. The primary goal is to develop a theoretical framework for the analysis of single-molecule two-color and three-color fluorescence data, which allows one to obtain biologically relevant information on biomacromolecules. Another goal is to understand the dynamics of complex biochemical reactions such as multisite enzyme catalysis and multisite ligand binding.
Current Research
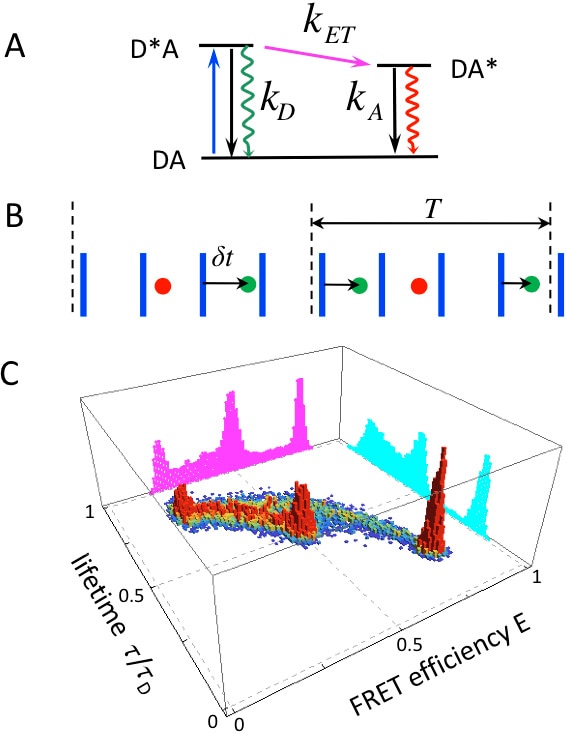
My current research is focused on two different topics. The first direction is related to single-molecule Förster resonance energy transfer (FRET) between fluorescent donor and acceptor labels attached to a biomacromolecule. Single-molecule FRET is widely used to probe intramolecular distances to study the structure, dynamics, and function of macromolecules. The output of these experiments is a sequence of photons with different colors and arrival times. Quantitative analysis of photon sequences requires a complete understanding of all microscopic processes involved. Extracting information about the distance fluctuations is particularly challenging when the fluctuations are as fast as the time between photons. In three-color FRET, three labels are attached to a molecule of interest (one donor and two acceptors). These experiments are much more difficult to interpret than the two-color FRET experiments. We develop a rigorous theoretical framework for the analysis of such raw two-color and three-color data. In collaboration with the leading groups of experimentalists studying various single biomacromolecules, we develop new methods to analyze single-molecule experiments and to obtain a microscopic picture of the specific biological system under investigation.
Another direction of our research is to understand the role of diffusion in the kinetics of a complex networks of bimolecular reactions such as repeated phosphorylation of a substrate by an enzyme or ligand binding to a macromolecule with several sites. We discovered that in general it is not enough to replace the rate constants of chemical kinetics by its diffusion-influenced counterparts, but one has to actually change the kinetic scheme by introducing new reaction channels between the bound complexes. This happens because a ligand that just dissociated from one site can diffuse to the other and then bind without diffusing into the bulk. We develop a general theory to treat such networks of diffusion-influenced reactions. The theory is applied to study multisite ligand binding, multisite phosphorylation, and stochastically gated reactions.
Applying our Research
The results of our fundamental research are the basis for the future development of precise tools to influence protein structure and the mechanisms of complex networks of biochemical reactions. These future tools are indispensable for applied science dealing directly with the problems of alleviation of a plethora of human adverse health conditions.
Select Publications
- Maximum Likelihood Analysis of Diffusing Molecules with Conformational Dynamics in Single-Molecule FRET.
- Gopich IV, Louis JM, Chung HS.
- J Phys Chem B (2025 Feb 27) 129:2187-2200. Abstract/Full Text
- Unraveling Burst Selection Bias in Single-Molecule FRET of Species with Unequal Brightness and Diffusivity.
- Gopich IV, Chung HS.
- J Phys Chem B (2024 Jun 13) 128:5576-5589. Abstract/Full Text
Research in Plain Language
To create new drugs to fight a disease, scientists need to know the structure and function of biological macromolecules, which are the basis of our life. Innovative techniques allow to observe the behavior of a single molecule by looking at the light pulses coming from the molecule. However, to understand the information content of such light pulses, one needs a mathematical description of these experiments. Our theory gives researchers a tool to analyze such single-molecule data. Our theoretical descriptions also extend to understanding how different biological molecules interact with each other in our body. Such interactions are overly complex since they involve many coupled biochemical reactions. Understanding the mechanisms of such intricate processes will allow scientists to control these reactions and help to find a cure for associated disorders.
Research Images