About Our Research
The Gene Regulation and Development Section is working to understand how transcriptional enhancers activate gene expression during development. Enhancers are highly cell type specific and this activation underlies how cells and tissues become different from one another and eventually form a complete organism. Enhancers are typically distant from their target genes along the linear sequence of chromosomes and one focus in the lab is on how the genome is organized in the nucleus to bring enhancers and their proper gene targets into proximity with each other. Our mechanistic studies involve genetics, biochemistry, molecular biology and computational analysis of genome wide data sets. Our studies contribute to understanding how differentiation and development are organized at the level of gene transcription.
Current Research
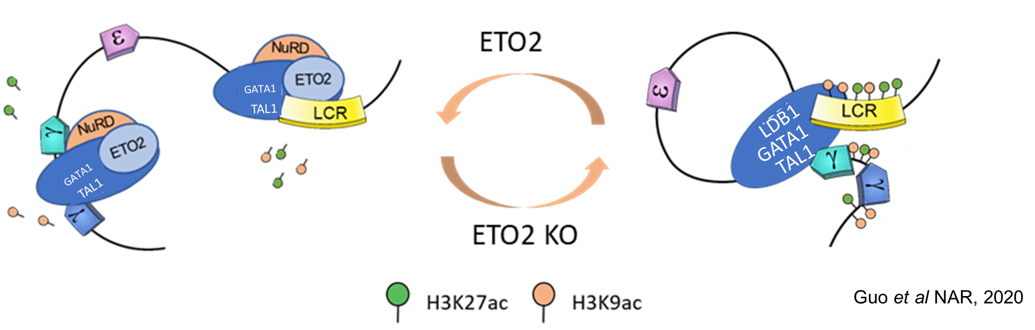
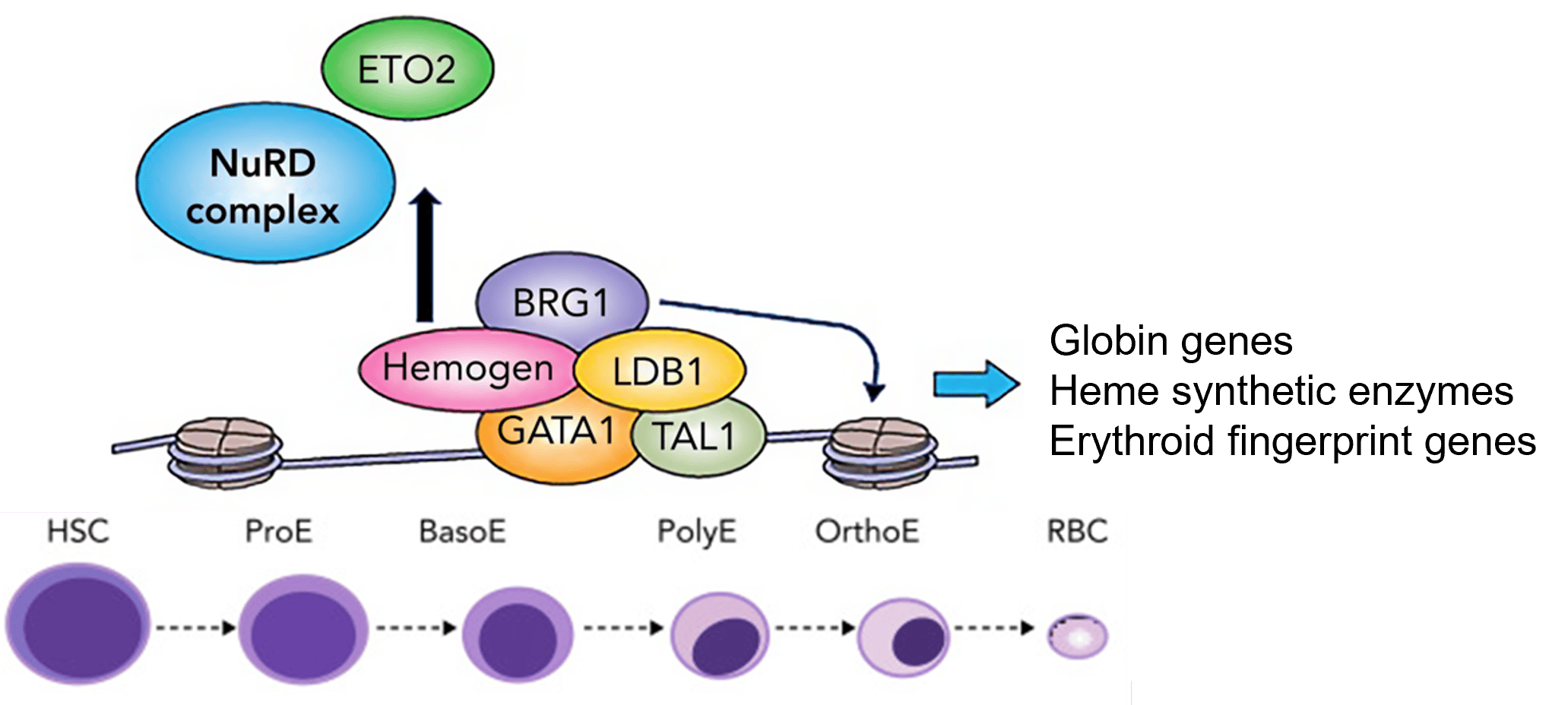
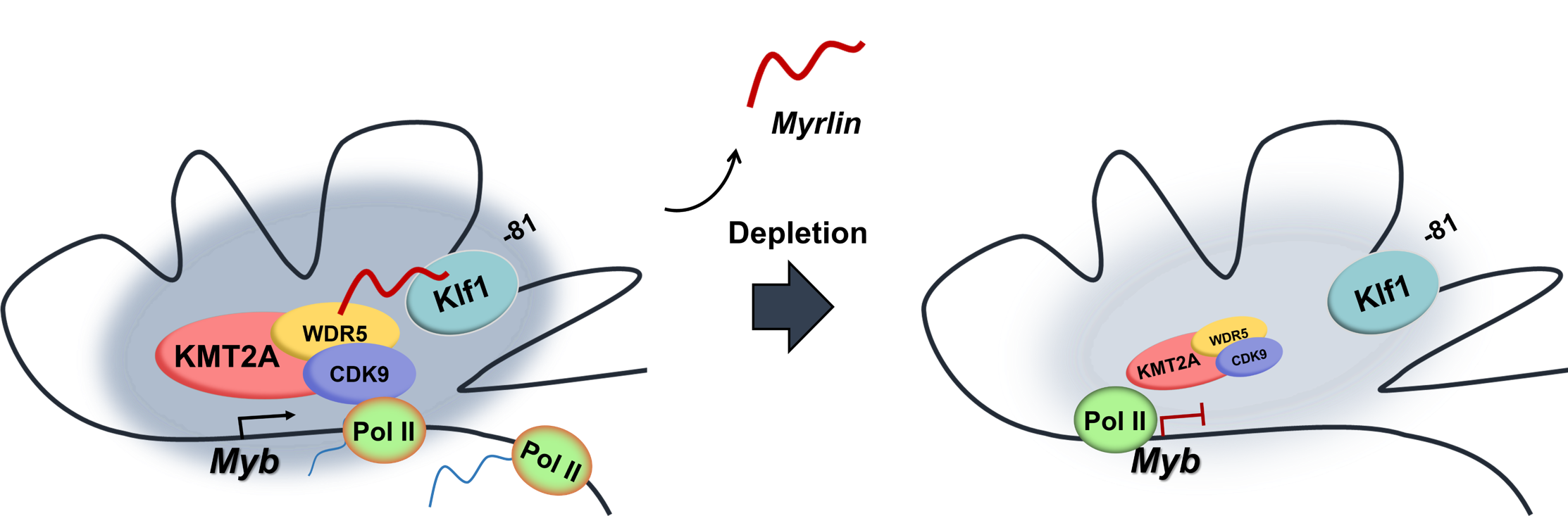
Our research is focused on how genes are turned on and off in developing cells of the hematopoietic system that is responsible for cells in the blood. The mechanisms of gene expression that we are trying to understand using the globin genes and other erythroid-cell-expressed genes are a paradigm for gene regulation in many systems.
We seek to understand how enhancers function at a fundamental level to deepen our understanding of how gene expression guides development and differentiation. The ultimate goal is to apply this knowledge to genetic diseases of red blood cells such as sickle cell disease and beta-thalassemia in which the hemoglobin molecule (that carries oxygen in the blood) is abnormal either in structure or amount.
Research that is ongoing in our lab and others suggest that it may be possible to manipulate the activity of enhancers, either positively or negatively, to ameliorate genetic diseases of the hematopoietic and other system and cancers.
Research Images