About Our Research
The Genetics of Organelle Biogenesis Section studies the biogenesis and function of centrosomes, non-membrane-bound organelles that organize the microtubule cytoskeleton into large macromolecular machines such as the mitotic spindle and cilia.
We seek to understand the molecular mechanisms that govern the number and size of centrosomes. These non-membranes bound organelles play a central role in cell growth, division, signaling, and motility and are present at one to two copies per cell. Although defects in centrosome structure and number can have catastrophic effects on cell division and genome integrity, much remains to be learned about how the number and assembly of centrosomes is controlled.
Our current research focusing on identifying and characterizing genes required for assembly of the two components of the centrosome; the centrioles and the pericentriolar material. Our long-term goal is to define in molecular term, the regulatory networks that govern the number and size of centrosomes.
Our lab wants to understand how cells control centrosome number and size as these parameters are critically important for proper cell growth, and division. To address these questions, we use the small nematode worm C. elegans as a simple animal model system. Most worm genes have human counterparts and thus our identification of genes that control centrosome number and size in the worm can inform us about analogous processes that operate in humans. During the last decade, it has become increasingly apparent that defects in centrosome number and size are associated with a variety of human diseases and thus a more complete understanding of how the centrosome is regulated will allow us to better understand how diseases arise as a result of centrosome dysfunction.
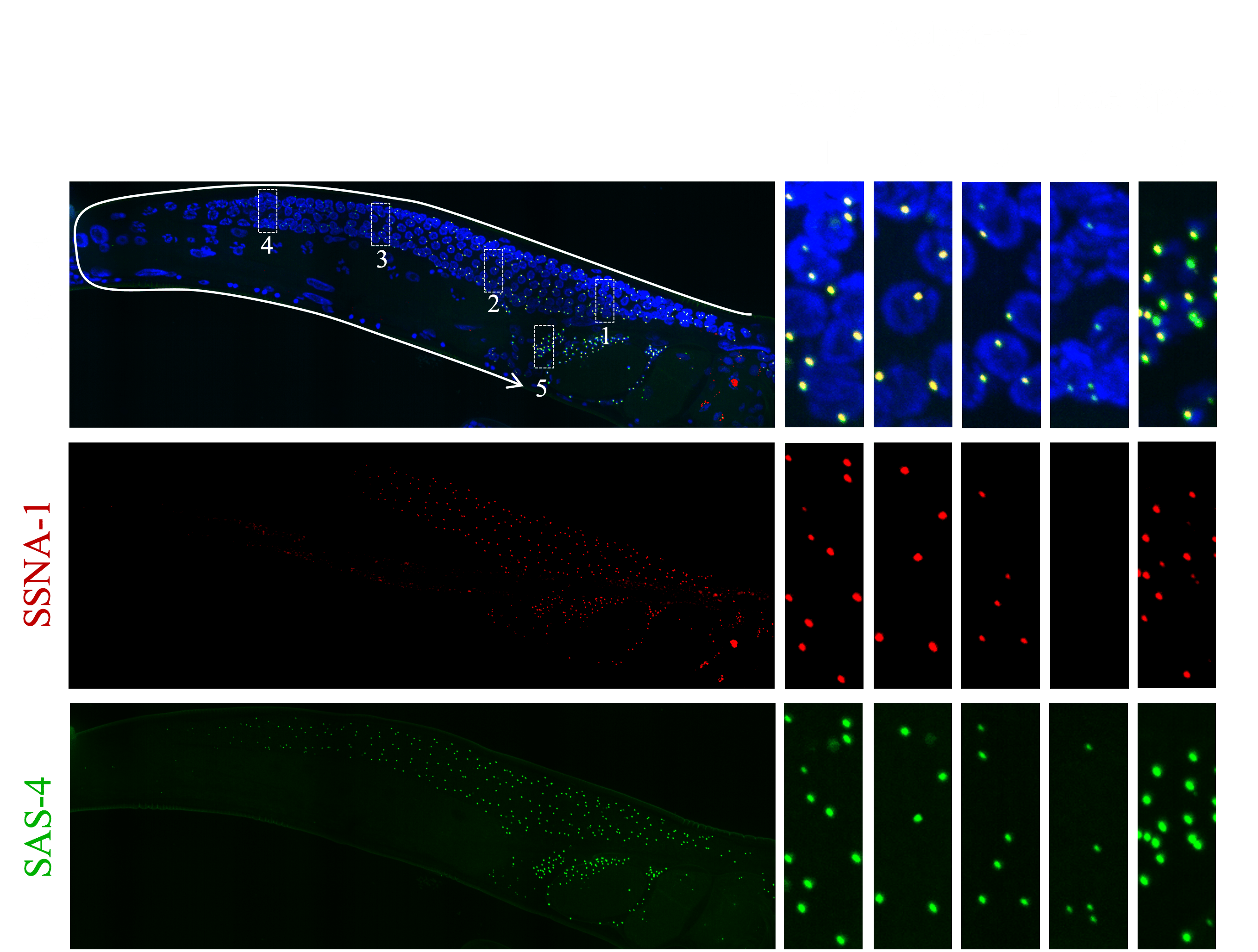
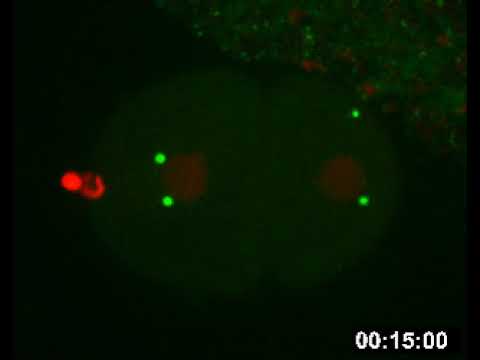
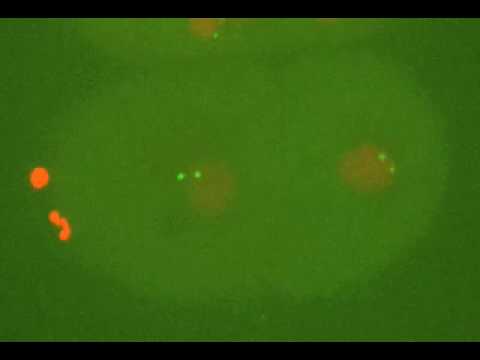
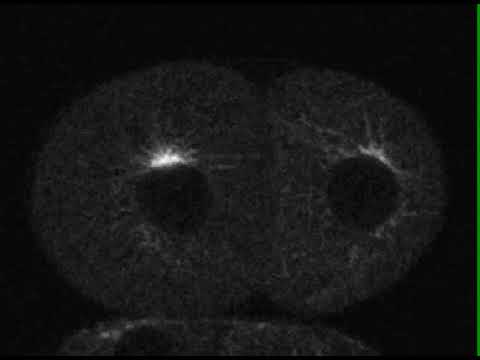
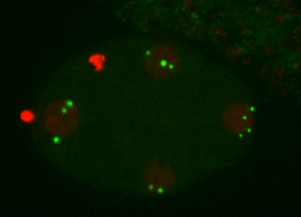
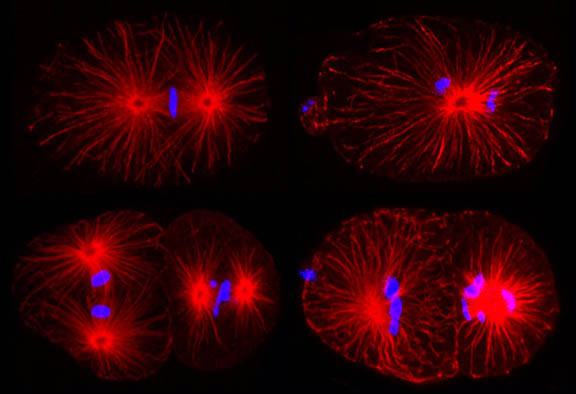
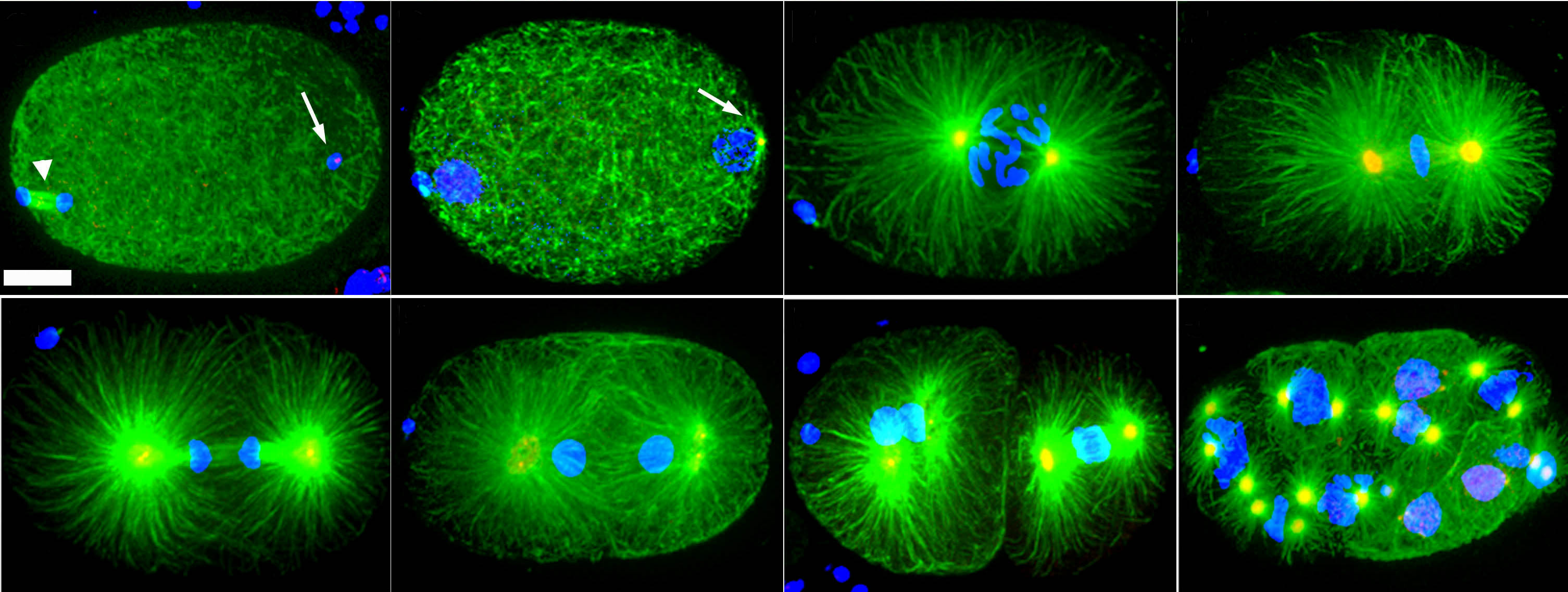
Research Images