About Our Research
We study conformational dynamics of proteins using single molecule Förster resonance energy transfer (FRET) spectroscopy. Especially, we focus on intrinsically disordered proteins (IDPs) that are closely related to various human diseases. The primary goal of our research is to understand the mechanisms of binding and aggregation processes of IDPs.
Intrinsic Disordered Proteins and Human Disease
Proteins play critical roles in virtually every cellular process. To perform their biological functions, they must properly fold from a disordered state into a unique three-dimensional structure. Bioinformatics studies, however, have revealed that a surprisingly large fraction (> 30%) of eukaryotic proteins actually contain a long stretch of disordered regions. These unstructured proteins or regions of polypeptides are called intrinsically disordered proteins (IDPs) or intrinsically disordered regions (IDRs). Although the prevalence of IDPs and IDRs apparently contradicts the concept of structure-function relationship, these unstructured proteins usually fold when they attach to the binding partners. The advantage of binding mediated by IDPs and IDRs over interactions between folded proteins is that the disorder allows flexible binding to many different binding partners. Therefore, intrinsic disorder is found in many proteins performing central roles in cellular regulation such as gene transcription and signal transduction and is frequently implicated in the development of various diseases such as cancer. In addition, disordered proteins can induce the formation of membraneless organelles through liquid-liquid phase separation and are often prone to aggregation, which also causes various protein misfolding diseases.
Projects
Our research program consists of new experimental approaches towards understanding the fundamental mechanism of these two main aspects of the disordered protein dynamics, binding and aggregation, using single molecule Förster Resonance Energy Transfer (FRET) spectroscopy and fluorescence imaging.
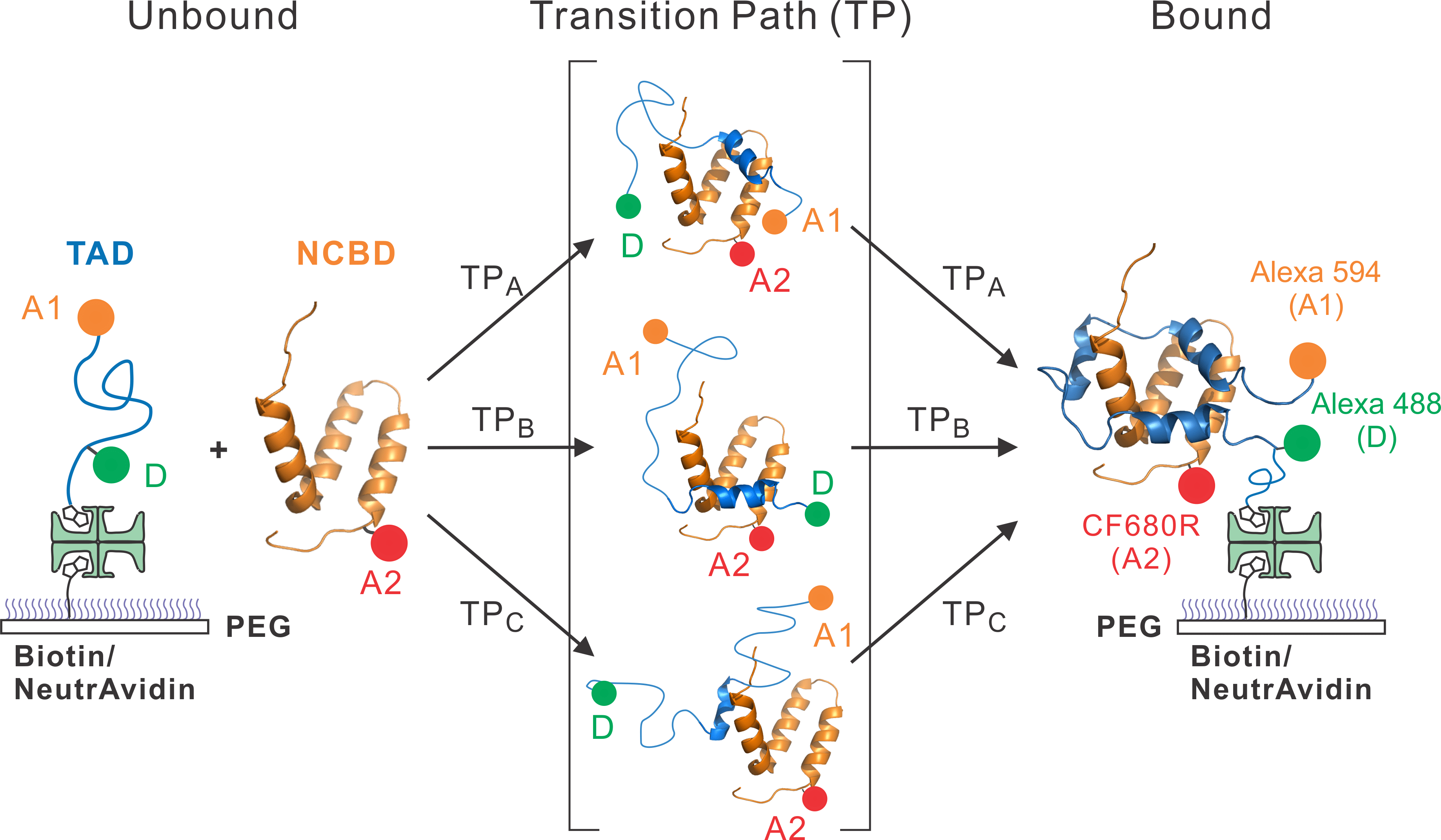
1. Understanding binding mechanism of IDPs. We try to understand the mechanistic details of binding such as how molecular conformations evolve when two molecules approach, make a contact, and form a bound complex. This information is contained in the moment of binding that can be probed only by analyzing individual events.
2. Characterization of protein aggregation. Protein aggregates and oligomers are thought to be implicated in the development of various neurodegenerative diseases. However, oligomerization and aggregation have been extremely difficult to study due to the heterogeneity of the process. Single molecule spectroscopy and fluorescence imaging can effectively characterize this complicated process by detecting individual molecular species without separation.
Tool: Fast Single Molecule FRET Spectroscopy and Fluorescence Imaging with Deep Learning
Recent development of various single-molecule techniques has allowed monitoring heterogeneous biological processes. Studying the heterogeneity is particularly important for understanding binding and aggregation mechanisms because conformational transitions of proteins are intrinsically heterogeneous.
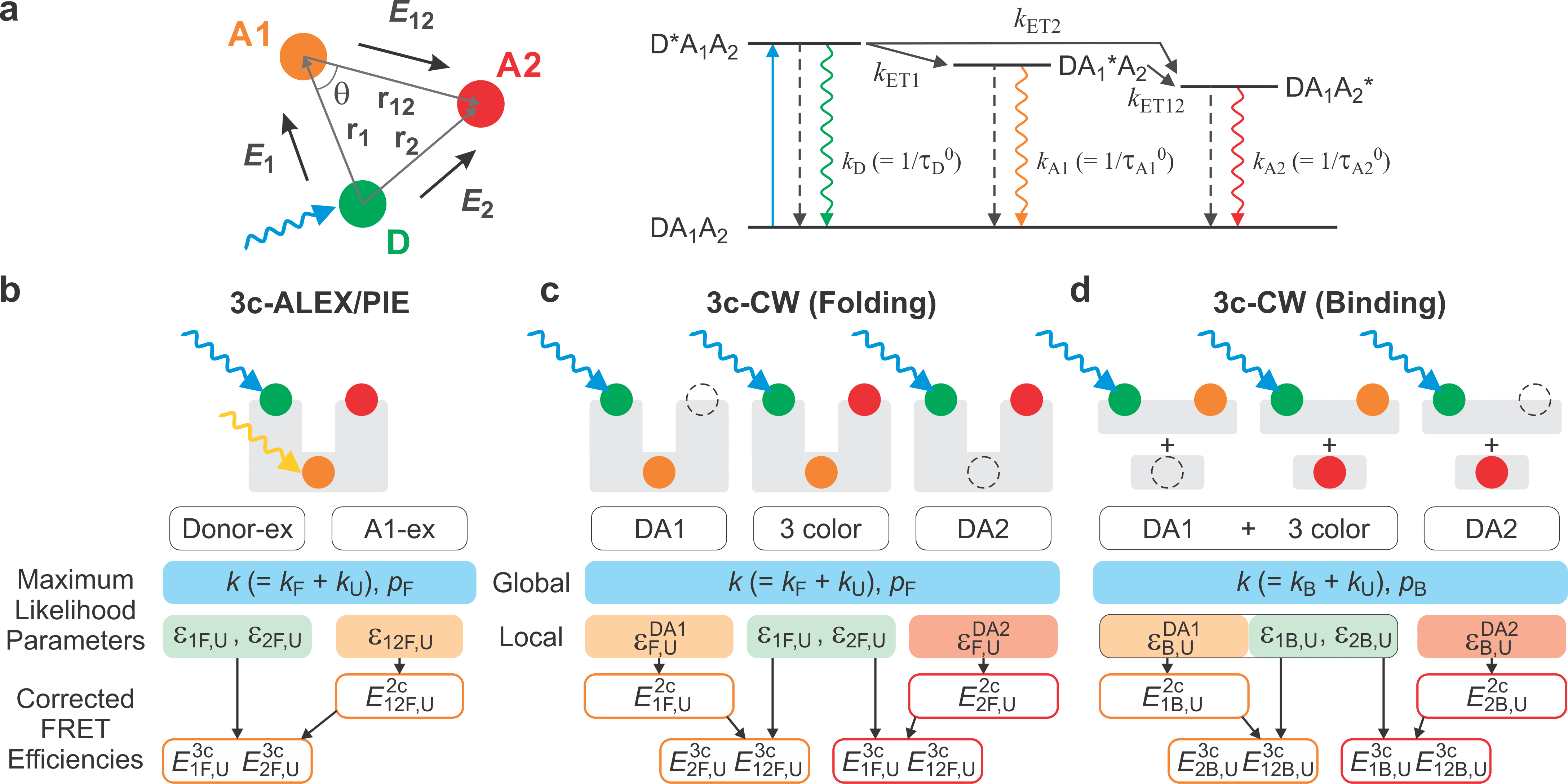
In FRET spectroscopy, the energy transfer efficiency depends strongly on the distance between a donor and an acceptor attached to a protein molecule. We are developing and using multi-color FRET spectroscopy to detect protein conformational changes of IDPs when binding happens. In order to capture fast binding events, we use a maximum likelihood method that analyzes information from individual photons.
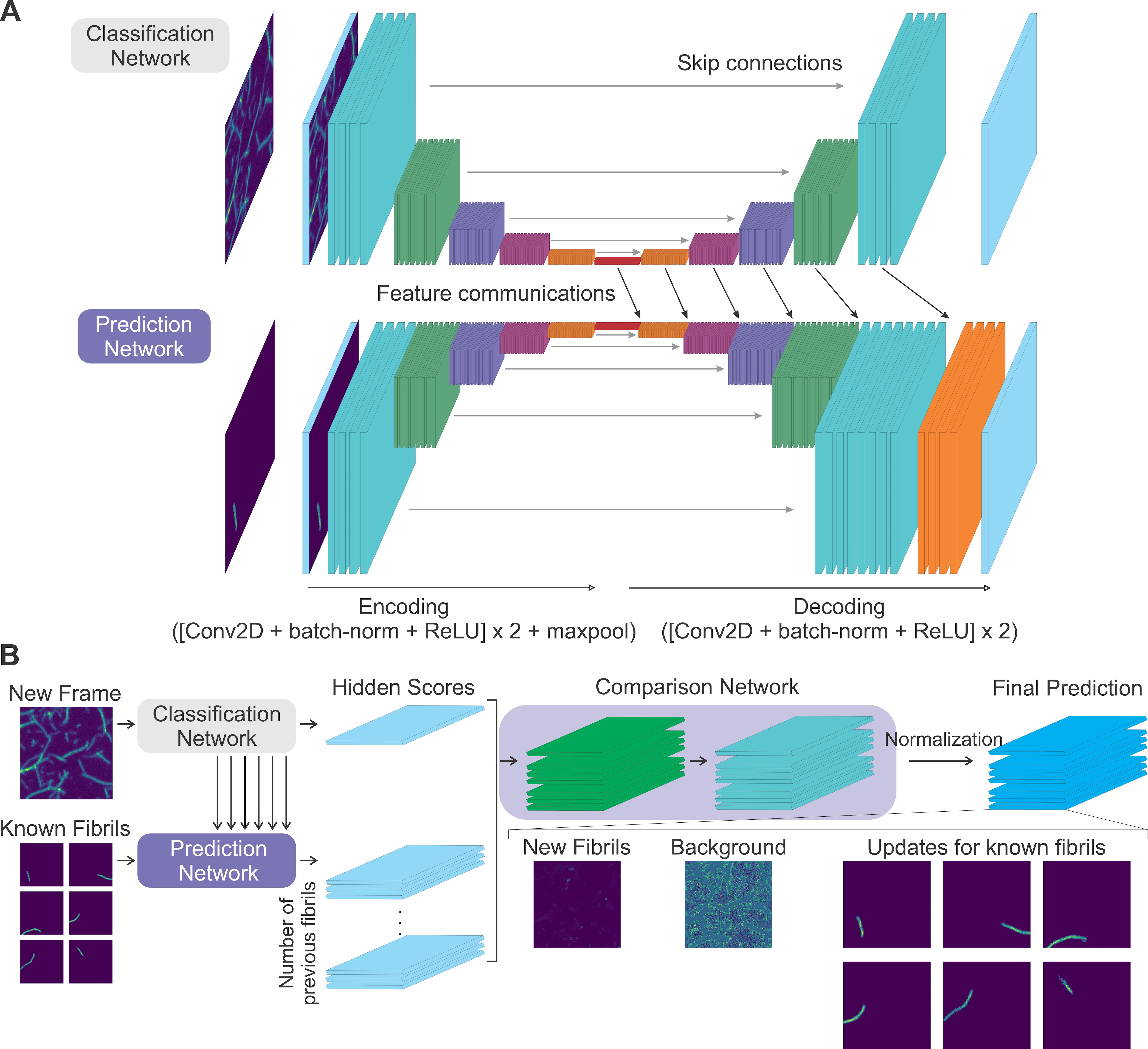
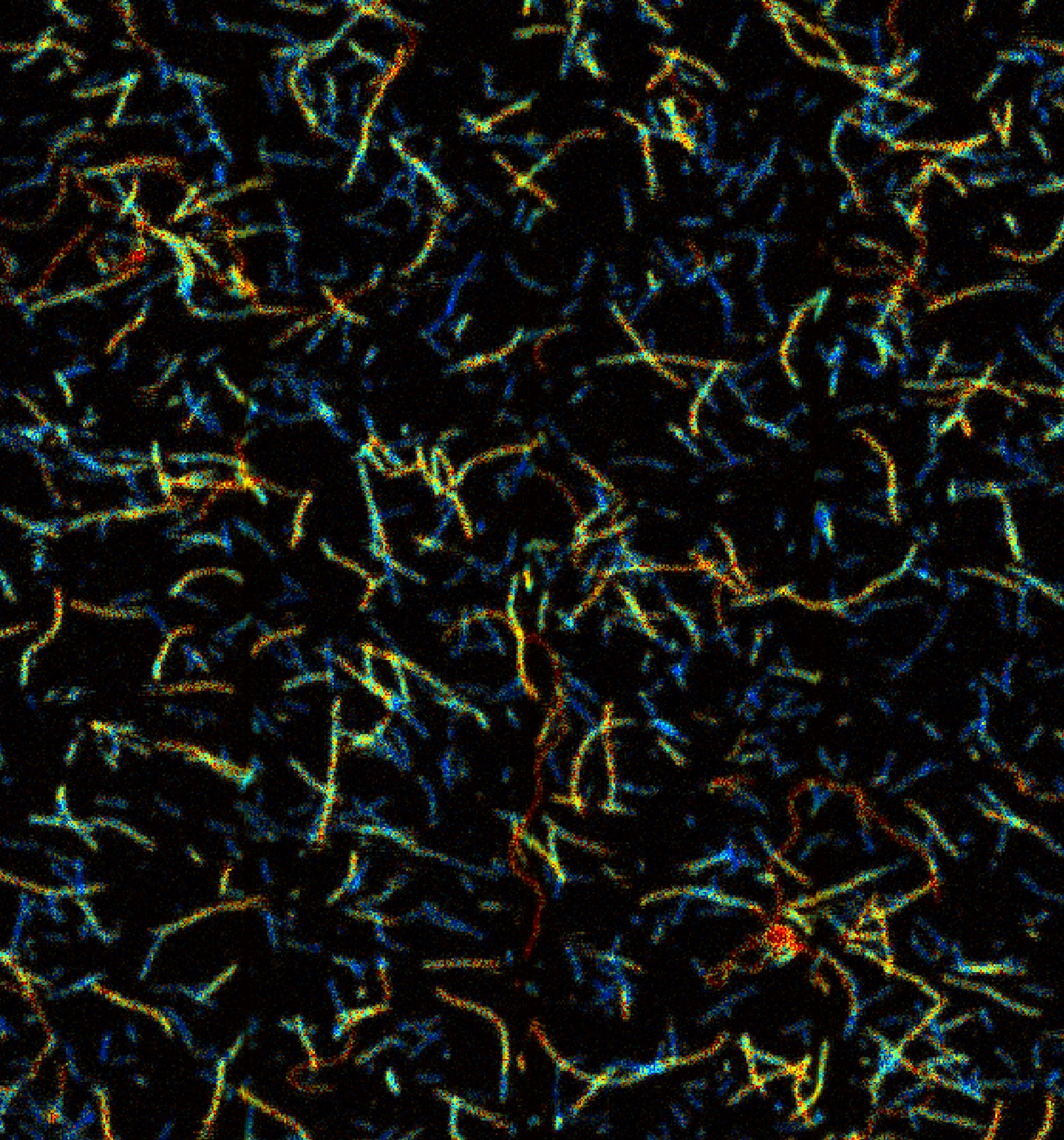
Characterizing heterogeneity in protein aggregation requires tracking the formation of individual amyloid fibrils. To this end, we are developing fluorescence imaging and deep learning based image segmentation methods for the analysis of single fibrils.
Research Images