Ad Bax Group Software
The Ad Bax Group makes available downloadable software for nuclear magnetic resonance (NMR) research.
Our software is available behind our new, NIH-required login.
Multiple login options—including Google and Microsoft—are freely available.

Available Software
- NMRPipe: Multidimensional spectral processing and analysis of NMR data
- TALOS: Prediction of Protein Phi and Psi Angles Using a Chemical Shift Database
- ACME: Measurement of homonuclear proton couplings from regular 2D COSY spectra
- SSIA: Simulation of Sterically Induced Alignment Tensor
- PALES: Prediction of ALignmEnt from Structure
- EHM: Extended Histogram Method for Analysis of Dipolar Couplings
- HBDB: Database Hydrogen-Bonding Potential for Protein Structure Refinement
- SAXS: Refinement of Protein Structures Against Small-Angle X-Ray Scattering Data
- SPARTA: Prediction of Backbone Chemical Shifts from Known Protein Structure
- CS-ROSETTA: Chemical Shifts Based Protein Structure Prediction Using ROSETTA
- IDIDC: Iterative DIDC analysis of RDCs
- FastSAXS: Fast refinement of macromolecular structures against solution x-ray scattering data
- TALOS+: A Hybrid method for predicting protein backbone torsion angles from NMR chemical shifts
- PROMEGA: Prediction of Xaa-Pro peptide bond conformation from sequence and chemical shifts
- SPARTA+: Improved Prediction of Backbone Chemical Shifts from Known Protein Structure
- MICS: Identification of Helix Capping and Beta-turn Motifs from NMR Chemical Shifts
- VW_fit: Optimization of weights of individual structural models in structural ensemble to achieve best fit for RDCs in multiple alignment media
- TALOS-N: Protein Backbone and Sidechain Torsion Angles Predicted from NMR Chemical Shifts Using Artificial Neural Networks
- POMONA: Chemical Shift Homology Modeling using Protein alignments Obtained by Matching Of NMR Assignments
- MERA: Backbone Torsion Angle Distributions Evaluation in Dynamic and Disordered Proteins from NMR Data
- SMILE: Sparse Multidimensional Iterative Lineshape-Enhanced (SMILE) Reconstruction of Both Non-Uniformly Sampled and Conventional NMR Data
- random coil J(HNHa): IDP random coil 3J(HNHa) coupling constants prediction
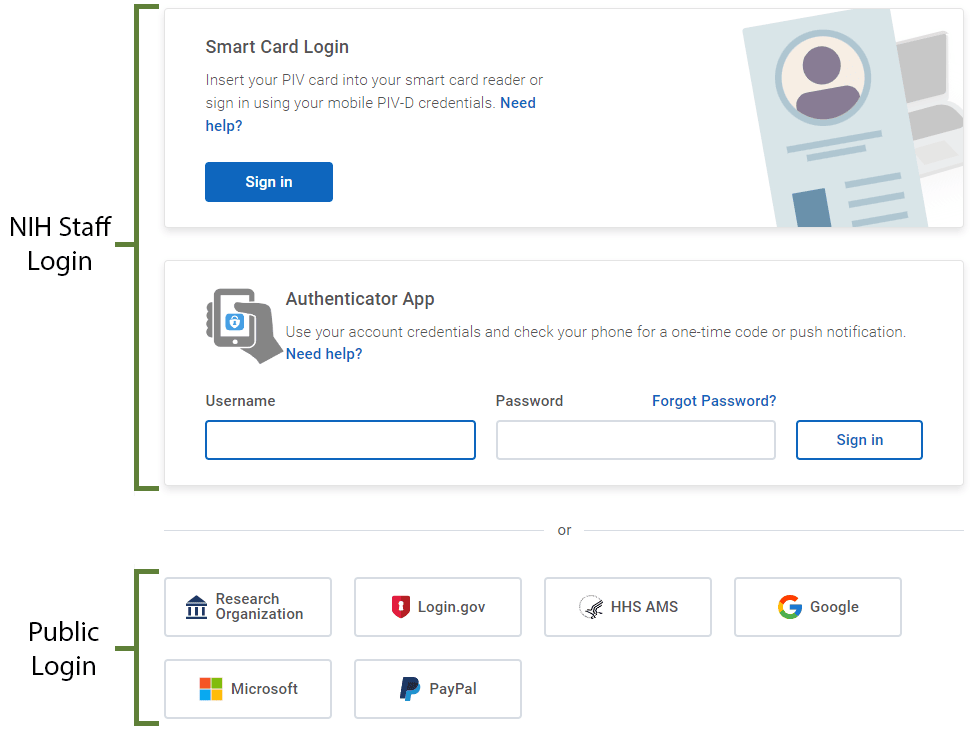
Use multiple login options—including Google, Microsoft, or NIH account—to access the Ad Bax Group software.
-
Click on the “Access Software” button.
-
NIH badge holders can use the “Smart Card Login” or “Authenticator App.”
-
Members of the public can scroll to the bottom of the page below the “Authenticator App” box and
- click on a preferred login option,
- follow the prompts to enter credentials, and
- confirm sharing account name with NIH.