Ad Bax Group Pulse Programs
The Ad Bax Group makes available pulse programs for nuclear magnetic resonance (NMR) research.
Our pulse programs are available behind our new, NIH-required login.
Multiple login options—including Google and Microsoft—are freely available.

Available Pulse Programs
- 2D IPAP [15N,1H] HSQC
- 2D IPAP HNCO
- 3D J-modulated HSQC
- 2D Constant-time TROSY J N-C'
- 3D TROSY HNCO J N-C'
- CBCA(CO)NH-JCH 3D Quantitative J Correlation
- 2D IPAP [13C,1H] CT-HSQC for Methyl Groups
- 3D Long Range 1H-1H Dipolar Coupling
- Long Range Dipolar Coupling with Nearest-Neighbor Decoupling
- Measurement of RDC in nucleic acid bases HC(C)-hd-TROSY-ECOSY
- Purine version
- Pyrimidine version
- H5C5-hd-TROSY Sample Data and NMRPipe Scripts
- H5C5(C6)-hd-TROSY-ECOSY Sample Data and NMRPipe Scripts
- Sample Data for intermediate processing and optimization
- Information on processing optimization
- 2D Selective CT 1H-31P NOESY
- 3D TROSY HNCO Quantitative J C'-CA
- 3D TROSY HN(CO)CA Quantitative J C'-CA
- Pulse Sequences for J(NC) in Nucleic Acids
- 3D spin-state selective HBCBCA
- 3D spin-state selective H5'C5'C4'
- 3D HNCA MQ Experiments for 3J Measurement
- 3D CT-MQ(1HN,13CA)+SQ(1HN)-HNCA spectra for 3JHN,HA (pulse code)
- 3D MQ(1HN,13CA)-HNCA experiments for 3JHN,CB (pulse code)
- 3D CT-MQ(1HN,13CA)-HNCA spectrum for 3JHN,C' (pulse code)
- 2D ARTSY for 1DNH RDC measurement
- CH2-TROSY HACAN
- Hydrogen Exchange (WEX-III) TROSY
- imino 15N-1H and base 13C-1H ARTSY for 1DNH and 1DCH RDC measurement in RNA
- SJS-HSQC for small molecule RDC (Angew. Chem. 2011, 50, 1-6)
- ARTSY-HNCO for measuring 1DHN couplings (JACS 2021, 143, 19306-19310)
- TATER-HNCO for measuring 2D(HN-C') couplings (JACS 2021, 143, 19306-19310)
- Anisotropic Shift Measurements:
- H1C1C2
- H1C1(C2)C3
- H1C1(C2C3)C4
- (H5)C5(C4C3C2)C1H1
- H1'->C1' -> C2' Out-and-Back CCH COSY
- CH2-TROSY Experiments:
- 2D [13C,1H] Correlation for DNA
- 2D [13C,1H] Correlation for Glycine
- 2D [13C,1H] Correlation for Protein Sidechains
- 3D [13C,1H,1H] TROSY-NOESY for DNA
- 3D constant-time parallel evolution HMQC-IPAP-NOESY
- 3D mixed-time parallel evolution HMQC-NOESY
- HSQC and TROSY experiments for measuring RNA base 13C residual CSA and pseudo-CSA shift
- 15N Relaxation Experiments (TROSY and sensitivity-enhanced HSQC)
- 1H T2 measurement by 2D Hahn-echo TROSY
- Homonuclear decoupling for enhancing resolution and sensitivity in NOE and RDC measurements of peptides and proteins
- Bruker acqu parameters and pulse code for 1D BASH (analog mode)
- Updated Bruker pulse code for 1D BASH in the digital mode
- Bruker acqu parameters and pulse code for 2D noesy with BASH
- Bruker acqu parameters and pulse code for HSQC with BASH
- Bruker acqu parameters and pulse code for 3D HA coupled hncoca with BASH
- HNCOCONH for the measurement of 3J(C'C')
- Accurate Measurement of (3)JHNHa Couplings in Small or Disordered Proteins from WATERGATE-optimized TROSY Spectra
- ARTSY-J: Convenient and precise measurement of 3JHNHa couplings in mediume-size proteins from TROSY-HSQC spectra
- Include File (includes definitions of items used in the pulse program)
Use multiple login options—including Google, Microsoft, or NIH account—to access the Ad Bax Group pulse programs.
-
Click on the “Access Pulse Programs” button.
-
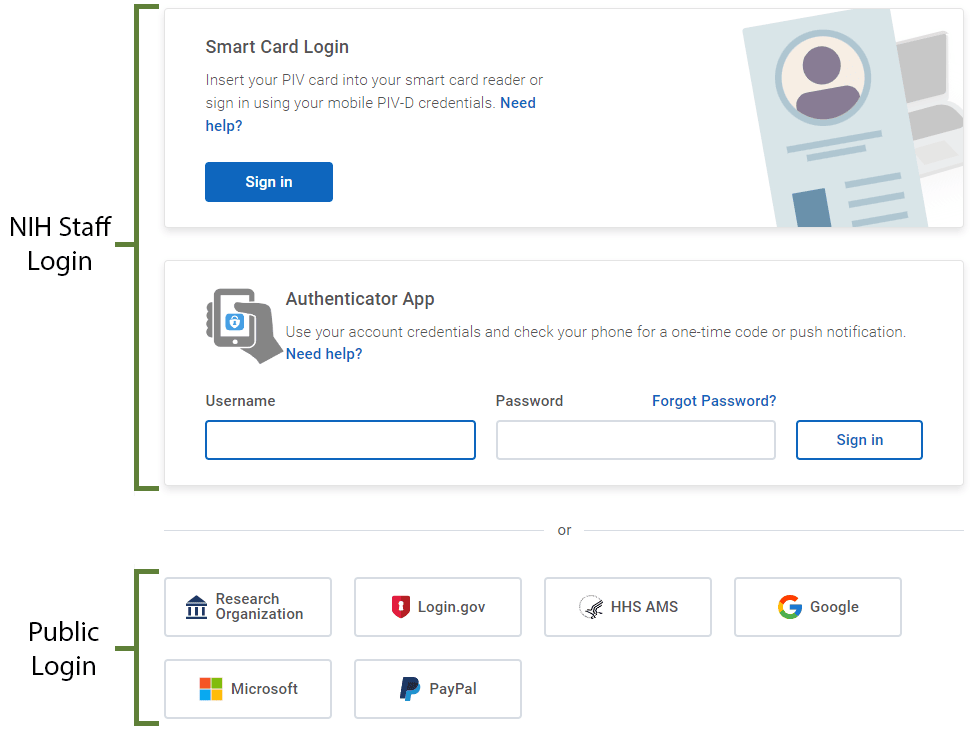
NIH badge holders can use the “Smart Card Login” or “Authenticator App.”
-
Members of the public can scroll to the bottom of the page below the “Authenticator App” box and
- click on a preferred login option,
- follow the prompts to enter credentials, and
- confirm sharing account name with NIH.